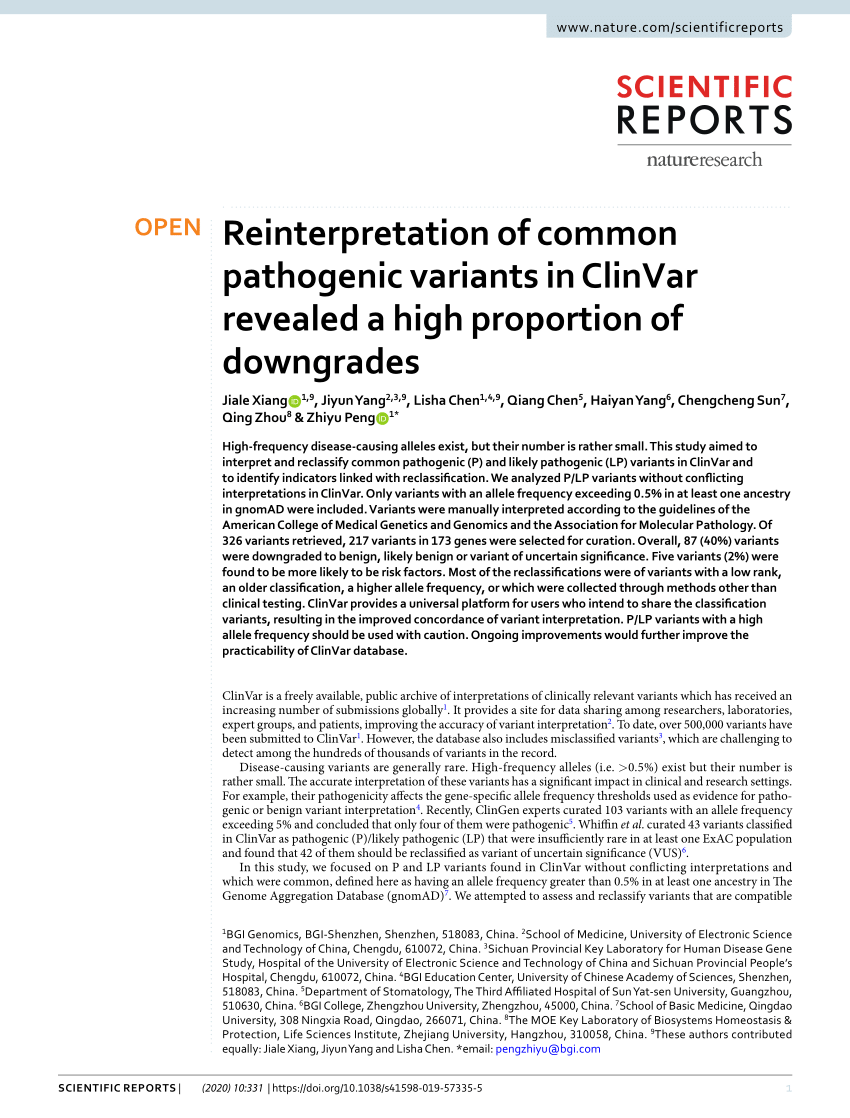

PDF) Reinterpretation of common pathogenic variants in ClinVar revealed a high proportion of downgrades

Clinical laboratories collaborate to resolve differences in variant interpretations submitted to ClinVar | Genetics in Medicine

CIViC is a community knowledgebase for expert crowdsourcing the clinical interpretation of variants in cancer | Nature Genetics
![PDF] ClinVar: improving access to variant interpretations and supporting evidence | Semantic Scholar PDF] ClinVar: improving access to variant interpretations and supporting evidence | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/128ba335c95595ea6846fed1c349601e97a18e5e/3-Figure3-1.png)
PDF] ClinVar: improving access to variant interpretations and supporting evidence | Semantic Scholar
![PDF] ClinVar: improving access to variant interpretations and supporting evidence | Semantic Scholar PDF] ClinVar: improving access to variant interpretations and supporting evidence | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/128ba335c95595ea6846fed1c349601e97a18e5e/2-Figure1-1.png)
PDF] ClinVar: improving access to variant interpretations and supporting evidence | Semantic Scholar
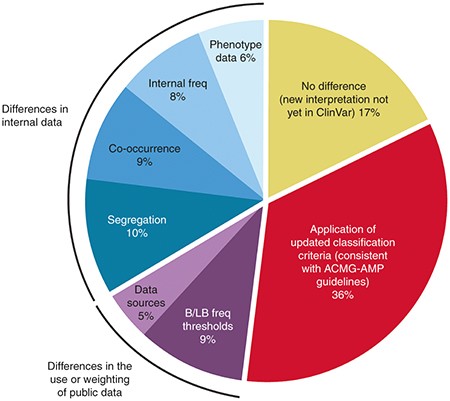
Clinical laboratories collaborate to resolve differences in variant interpretations submitted to ClinVar | Genetics in Medicine
![PDF] Clinotator: analyzing ClinVar variation reports to prioritize reclassification efforts | Semantic Scholar PDF] Clinotator: analyzing ClinVar variation reports to prioritize reclassification efforts | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a7c0e187c02501c23bc0ac6fa66512fd0cb1c655/7-Figure1-1.png)
PDF] Clinotator: analyzing ClinVar variation reports to prioritize reclassification efforts | Semantic Scholar
![PDF] ClinVar: improving access to variant interpretations and supporting evidence | Semantic Scholar PDF] ClinVar: improving access to variant interpretations and supporting evidence | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/128ba335c95595ea6846fed1c349601e97a18e5e/3-Figure2-1.png)
PDF] ClinVar: improving access to variant interpretations and supporting evidence | Semantic Scholar

Genome Alert!: a standardized procedure for genomic variant reinterpretation and automated genotype-phenotype reassessment in clinical routine | medRxiv
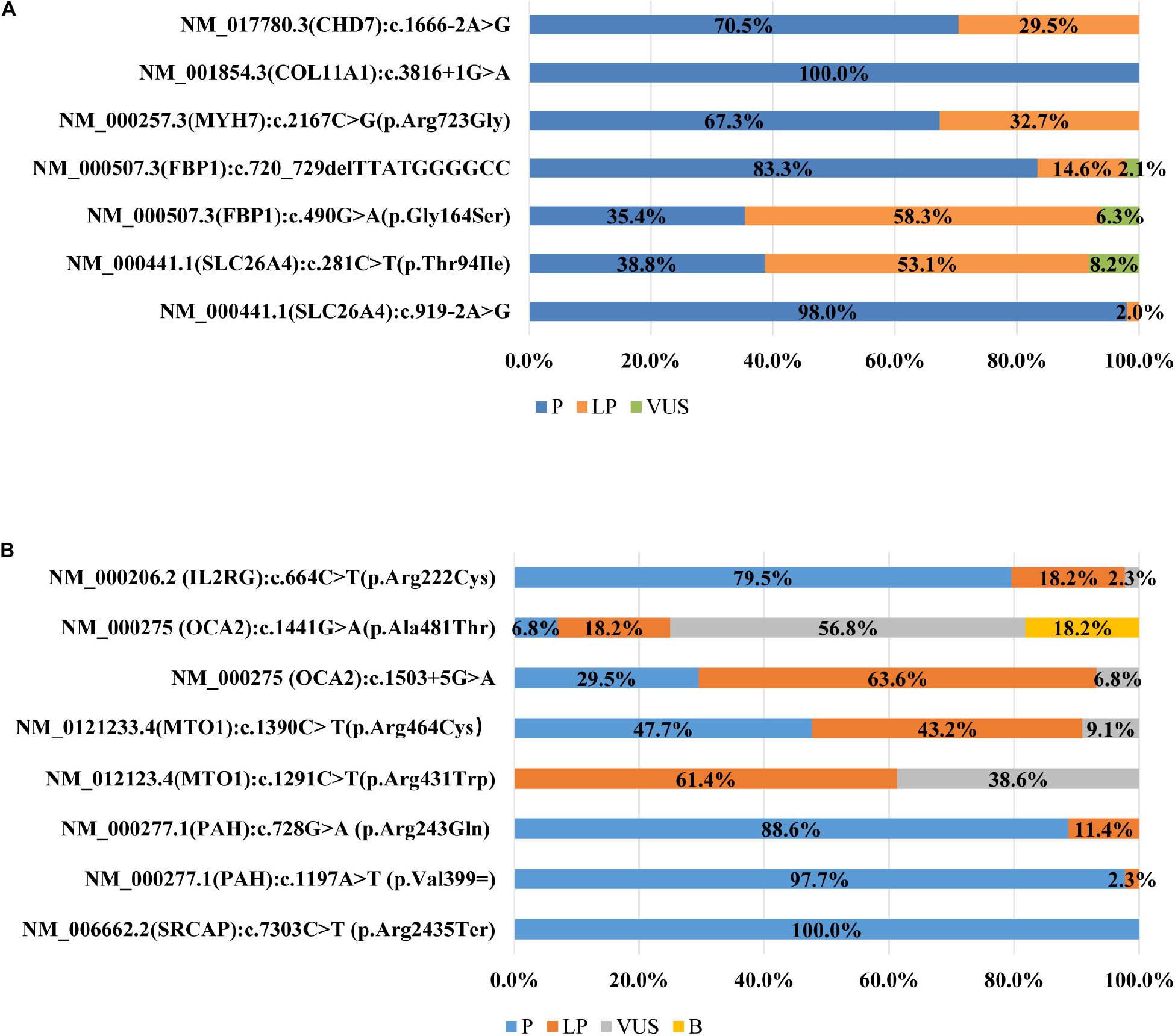
Frontiers | An Initial Survey of the Performances of Exome Variant Analysis and Clinical Reporting Among Diagnostic Laboratories in China

EVA/ClinVar - include other clinical significant variant · Issue #1139 · opentargets/issues · GitHub
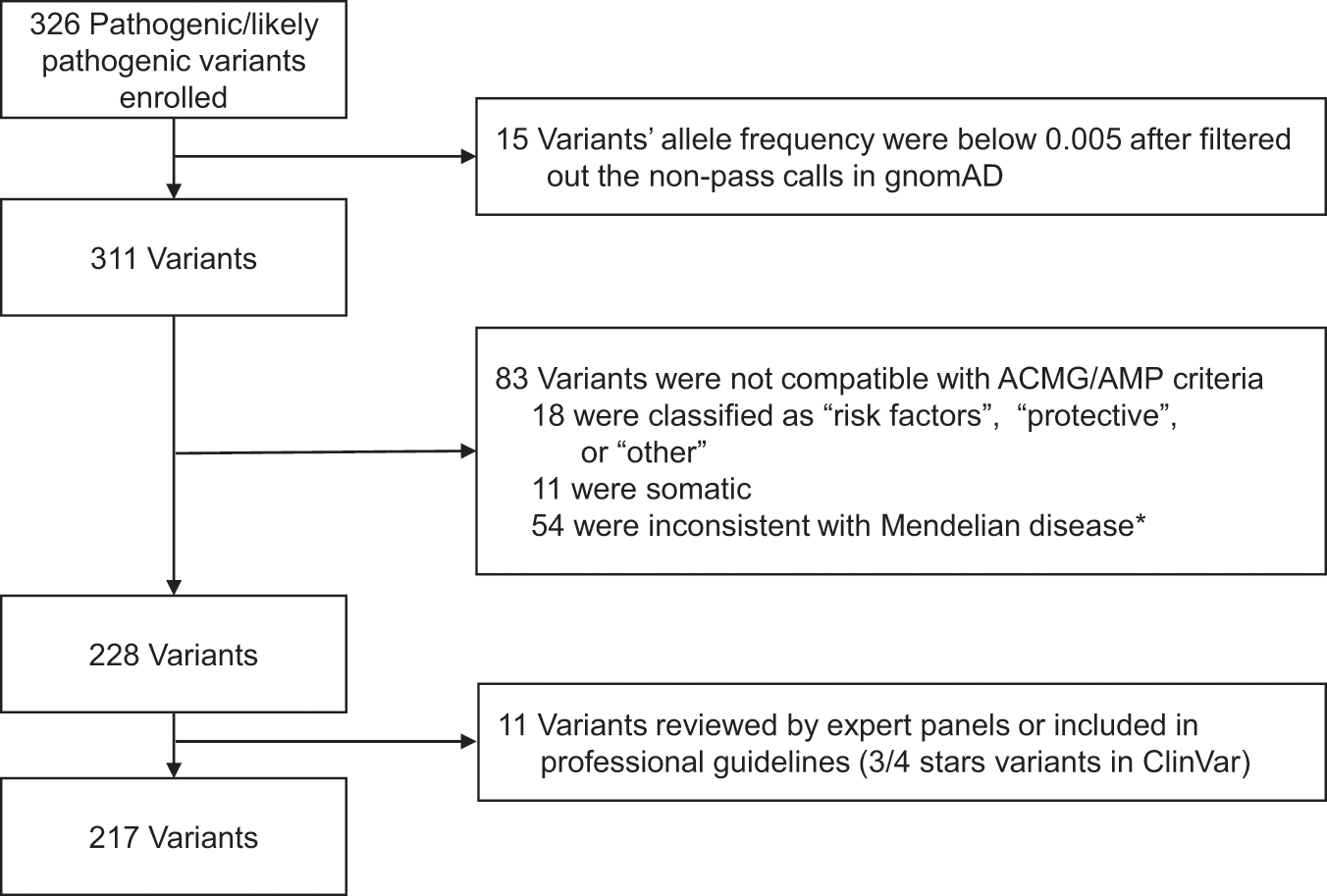

Reinterpretation of common pathogenic variants in ClinVar revealed a high proportion of downgrades | Scientific Reports








![PDF] ClinVar: public archive of interpretations of clinically relevant variants | Semantic Scholar PDF] ClinVar: public archive of interpretations of clinically relevant variants | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a77e1886e8d8659f89c6dfacbbf339cb3c5c0672/4-Figure2-1.png)



